Researchers often default to "the broader the panel, the better." But once your question narrows—say you only need a handful of adipokines across two visits—running large, disease‑area panels can become inefficient or misaligned. Two anonymized scenarios we see frequently illustrate the pivot point:
- Case A (running case throughout): A cardiometabolic study with 66 patients sampled at baseline and follow‑up. The team only needs 5–6 adipokines, yet those proteins are scattered across different Olink panels. Do they really need to run multiple broader panels to quantify a tiny, well‑defined set? Or is a small custom route more logical?
- A peptide‑vaccine oncology project wants a tight cytokine/chemokine subset to track vaccine response. The PI is weighing a standard inflammation panel against a semi‑custom panel or a deeper custom path tailored to a known signature.
Here's the deal: standard Olink panels remain excellent when your aim is broad pathway coverage or early hypothesis generation. But when targets are few, cross‑panel, or when you're verifying a specific signature, a custom biomarker set Olink approach—via Flex or Focus—may fit better. This guide shows where the boundary lies, then maps Flex (semi‑custom, library‑based) and Focus (signature‑oriented custom) to real project intents.
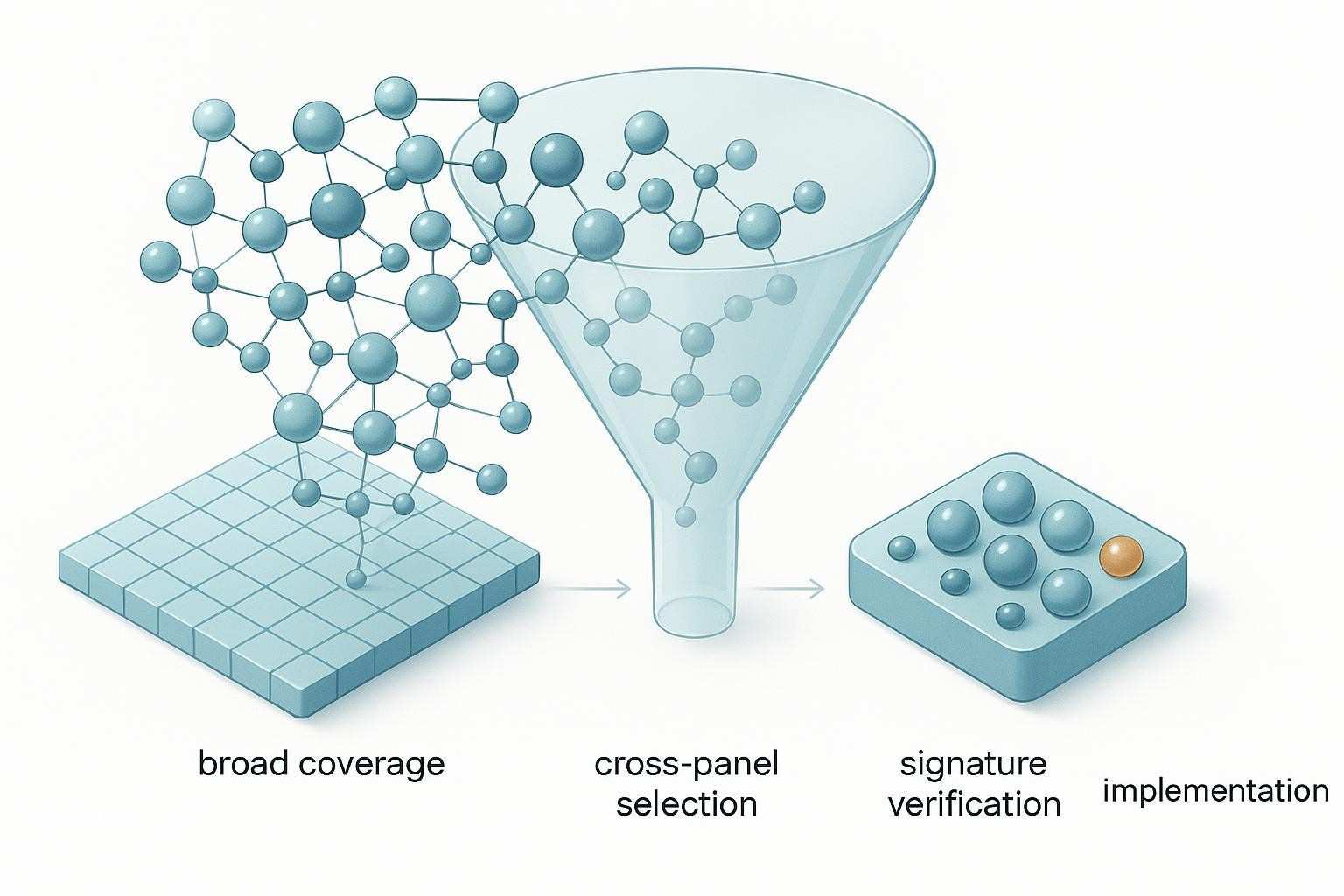 Figure 1. As research questions narrow from broad pathway coverage to defined biomarker signatures, the logic of panel selection shifts from standard multiplex panels toward more customized solutions.
Figure 1. As research questions narrow from broad pathway coverage to defined biomarker signatures, the logic of panel selection shifts from standard multiplex panels toward more customized solutions.
Standard panels are powerful—but they are not built for every narrow question
Standard Olink panels (e.g., disease‑area Target 96 or subsets within exploration workflows) are designed to cast a wide biological net. They shine in early discovery, when you want coherent pathway coverage, or when your indication maps cleanly to an existing, well‑validated panel. In translational projects, a standard panel often remains sufficient until the question tightens to a short list of proteins or a specific signature that doesn't align perfectly with a predefined panel.
When a standard panel is usually the right answer
- You need broad, hypothesis‑generating coverage across a pathway or disease area.
- Your focus area already aligns well with a standard disease‑area panel.
- The study is still exploratory; you're not constrained to a very small set.
- Cohort‑level profiling and pattern finding matter more than single‑marker precision right now.
For background on matching panels to broad study aims, see the neutral primer Choosing the Right Olink Panel from Creative Proteomics' knowledge hub: Choosing the Right Olink Panel.
The signs that a standard panel may be too broad, too rigid, or misaligned
- Only 5–10 proteins truly drive your analysis plan or decision endpoint.
- Your proteins of interest are scattered across multiple panels.
- You've moved from discovery to verification or implementation.
- Interpretability would improve by constraining to a small, well‑justified set.
- You're implementing a defined signature rather than exploring an entire biology area.
In Case A (66 patients; two time points; 5–6 adipokines), the team quickly realized that buying and running multiple broader panels would collect a lot of data they didn't plan to analyze—while still failing to deliver a tidy, focused readout aligned to their primary endpoints.
When standard Olink panels still make sense—and when they start to feel too broad
| Study situation | Standard panel fit | Why it may still work | Early sign you may need a custom route |
| Early discovery across a pathway (e.g., cardiometabolic) | Strong | Breadth helps surface hypotheses and networks | Only a handful of proteins consistently matter across analyses |
| Disease area maps neatly to a predefined panel | Strong | Validated, coherent coverage; easier interpretation | Key targets missing or split across panels |
| High‑level profiling for clustering/stratification | Moderate–Strong | Wider scope supports unsupervised findings | You already have a short, predefined set to verify |
| Longitudinal validation with a small target set | Weak–Moderate | Breadth becomes redundant and cost‑inefficient | You can name the few proteins that drive decisions |
The three most common situations where researchers start considering Flex or Focus
A common turning point appears when researchers realize the panel in hand is solving yesterday's question. Today's question is narrower and more actionable.
Scenario 1: You only care about a small number of proteins
Think 5–6 adipokines, a focused inflammation subset, or a small verification set from earlier discovery. In Case A, the cardiometabolic team needed leptin, adiponectin, resistin, and two or three adjunct markers. Collecting 80–90 additional proteins "just because they're on the panel" risked diluting attention and budget.
Scenario 2: Your targets are spread across multiple existing panels
Targets often live across inflammation, cardiometabolic, and immunology panels. Researchers don't want to run several broader panels just to capture a few scattered proteins. In practice, the most useful starting point is not the product label, but a list of proteins of interest with biological justification and matrix context.
Scenario 3: You are moving from discovery to targeted validation or implementation
The point is no longer "more data." It's whether a short signature survives focused testing and supports a translational decision. Teams frequently ask for a "small custom panel" before clarifying whether they're selecting known proteins from an existing library (a Flex‑type question) or implementing a defined signature with tailored validation (a Focus‑type question).
Embedded mini‑cases:
- Case A (primary, cardiometabolic): 66 patients; two time points; 5–6 adipokines → library check first (Flex) vs signature‑oriented development (Focus) if library gaps/validation needs arise.
- Case B (oncology peptide‑vaccine): Custom cytokine/chemokine set; the team explored a Focus 21 for a signature readout, plus a Flex 30 add‑on for additional immunology markers in a separate sub‑study.
- Case D (discovery→validation): After large‑scale discovery, only a narrow signature remained relevant for verification; this is a classic gate for Focus‑style implementation if the targets and context are well defined.
What Olink Flex is best suited for
A semi‑custom route built around a pre‑validated library
Olink Flex is designed for small, configurable panels by mixing and matching from a library of pre‑validated assays. Current Olink materials describe assembling approximately 5–30 biomarkers from ~200 assays, with options for minimal sample input (often 1 µL plasma/serum) and readouts in NPX by default; pg/mL may be available where Olink provides a calibrated route on compatible instrumentation (Olink Flex; Olink Signature Q100). Olink emphasizes high combinability and panel‑specific QC in this workflow (Olink Flex).
Where Flex shines in real project planning
Flex typically shines when your targets already exist in the library and you want a small, targeted panel without embarking on deeper custom co‑development. For teams starting from standard panels, Flex can be thought of as a "configure‑from‑library" path: select what you need, avoid what you don't, and keep the focus on the biology driving decisions. In Case A, if all 5–6 adipokines are in the library and matrix context is standard (e.g., plasma/serum), Flex is often the most efficient next step.
The practical limits researchers should understand
Flex is still bound to its library. Not every novel biomarker will be available, and "custom" here does not mean unconstrained assay invention. If you need to implement a defined signature with matrix‑ or context‑specific validation, or if library gaps are material, you may be outside Flex's sweet spot. Olink's documentation also cautions that NPX is the standard relative unit; absolute pg/mL requires assay‑ and instrument‑specific calibration and isn't universal (Olink FAQ).
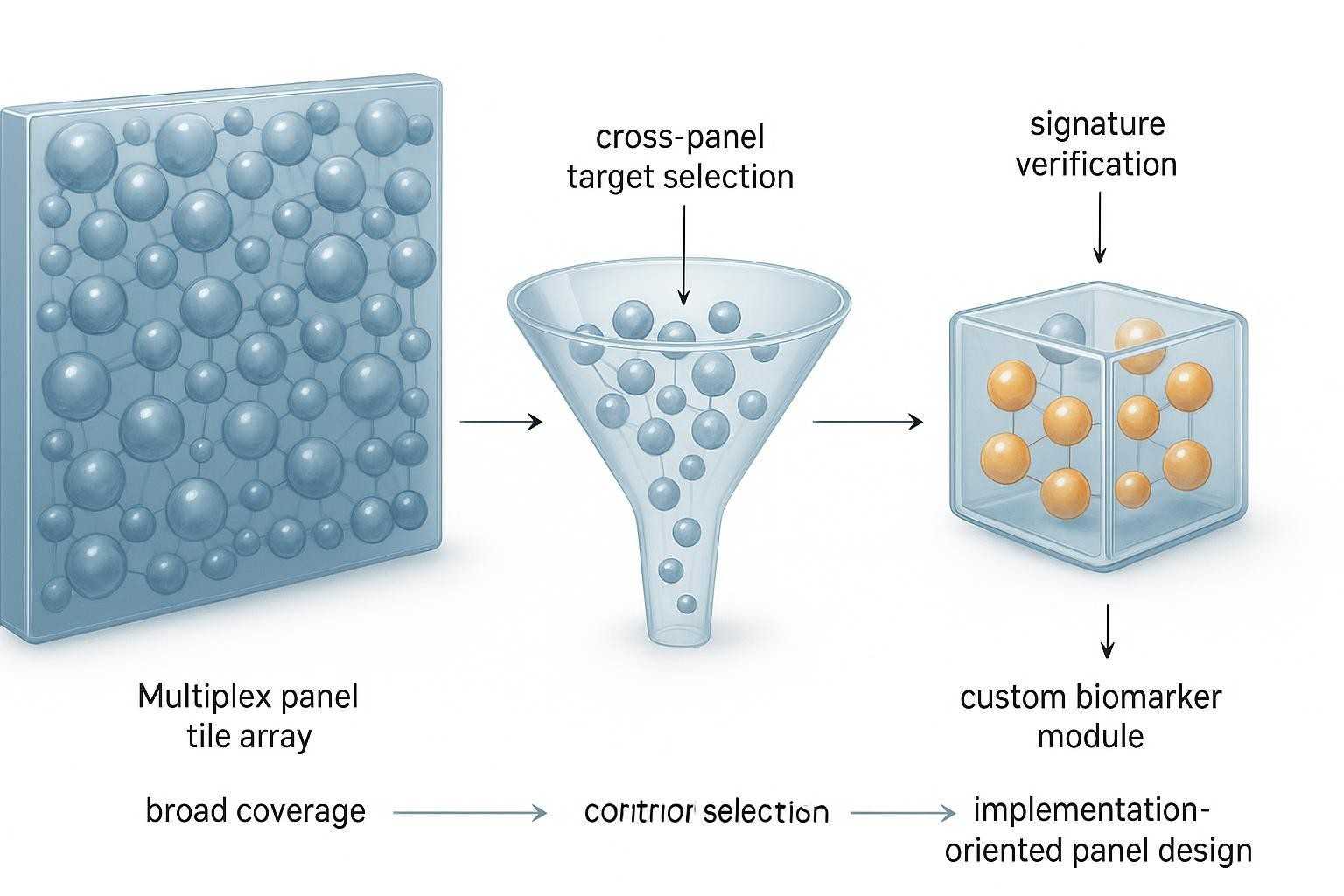 Figure 2. Flex and Focus occupy different parts of the custom biomarker workflow: one centers on configurable library‑based panels, while the other supports deeper signature‑oriented implementation.
Figure 2. Flex and Focus occupy different parts of the custom biomarker workflow: one centers on configurable library‑based panels, while the other supports deeper signature‑oriented implementation.
What Olink Focus is best suited for
A deeper custom route for implementing a protein signature
Olink Focus supports up to 21 proteins and taps a broader biomarker universe (>5,000 according to Olink), built through custom development by Olink's in‑house R&D experts. It's explicitly positioned for implementing or verifying biologically defined protein signatures, with NPX or (where available and appropriate) pg/mL outputs on compatible instruments (Olink Focus; Olink Signature Q100; Olink discovery→utility overview).
Where Focus becomes the more logical choice
Choose Focus when your target selection is more bespoke, your signature is biologically defined but not well served by standard panels, or when matrix‑/context‑specific validation matters more. This is common after discovery narrows to a compact signature and the study moves closer to translational decision‑making. In Case B (oncology peptide‑vaccine), a Focus 21 was used to lock the signature readout, while a separate Flex 30 module captured ancillary markers for exploratory immunology endpoints.
Why Focus should not be described as "just a smaller panel"
Focus is not merely fewer proteins. It reflects a different project intent (signature confirmation/implementation) and a different development model (tailored validation, broader sourcing). It sits closer to verification and actionability than to "trimmed‑down exploration."
Peer‑reviewed work underscores the reality behind these choices: PEA‑based panels typically report NPX as a relative, log2 unit; moving to absolute pg/mL requires validated calibration and isn't one‑size‑fits‑all (Carlyle 2022, Frontiers in Neurology; Heylen 2024; Siegbahn 2023).
Flex vs Focus is not the first question—this is
Before picking between Flex and Focus, step back. Are you sure a standard panel isn't already the most efficient answer? Then, if a custom path is warranted, ask whether you're selecting known proteins from an existing library or implementing a defined signature that needs tailored validation.
First ask: is a standard panel already sufficient?
If your question maps cleanly to an existing disease‑area panel and you still need broad coverage, a standard panel is typically the fastest, cleanest path. Don't over‑customize too early.
Then ask: are your targets already represented in an existing library?
If yes, and you only need a small set, Flex often fits: configure a panel of ~5–30 markers from ~200 assays and avoid the "too‑broad" problem (Olink Flex). If no, consider Focus or revisit whether targets can be substituted without compromising biology.
Finally ask: are you selecting known proteins, or implementing a signature?
Selecting a few known targets from a library is a Flex‑type decision. Implementing a constrained, biologically defined signature—especially with matrix‑specific requirements—points toward Focus.
A practical decision framework for choosing between a standard panel, Flex, and Focus
Start from your biological question, not from product names
In practice, the most useful starting point is the biological decision you're trying to make. What result would change your next step? That answer should dictate the panel type, size, and validation depth.
Define whether your bottleneck is coverage, specificity, or implementation
- Coverage bottleneck: choose standard panels for breadth.
- Specificity bottleneck around a few markers: consider Flex.
- Implementation bottleneck around a pre‑defined signature: consider Focus.
Clarify whether your project is still exploratory or already narrowing to action
Exploratory questions value breadth; action‑oriented questions value focus and reproducible, context‑appropriate validation.
Decision framework: Standard panel vs Flex vs Focus
| Decision factor | Standard panel | Olink Flex | Olink Focus |
| Typical protein count needed | 48–96+ depending on product | ~5–30 from library | Up to 21 (signature‑oriented) |
| Source of proteins | Predefined disease/pathway panels | Pre‑validated assay library (~200) | Broader biomarker universe (>5,000) |
| Best project stage | Early discovery; broad profiling | Targeted verification when assays exist | Signature verification/implementation |
| Best fit for cross‑panel target set | Limited if split across panels | Strong if targets in library | Strong when library gaps or bespoke validation needed |
| Validation depth | Panel‑level, standardized | Panel‑specific QC; NPX default | Tailored validation; NPX or pg/mL where calibrated |
| Typical researcher situation | Hypothesis generation; mapping pathways | Small known set; conserve budget/attention | Locking a defined signature for translational use |
| Main trade‑off | Breadth vs redundancy for narrow aims | Library‑bound; not all targets available | More development; focused scope |
Footnote: Flex and Focus specifications per Olink product pages (Olink Flex; Olink Focus). NPX is the standard relative unit; pg/mL availability is assay‑ and instrument‑dependent (Olink FAQ; Olink Signature Q100).
How to describe a custom biomarker request clearly when contacting a provider
In project discussions, teams often jump to product names ("Explore + Focus + Signature Q100 quote") before articulating the biology, targets, and study stage. A clearer request accelerates scoping and avoids mismatched expectations.
If you already have a shortlist of proteins
- Share the list with gene/protein names and any acceptable substitutes.
- Indicate species, matrix (e.g., plasma, serum, CSF), and expected concentration range if known.
- Note whether this is an exploratory shortlist or a locked signature for verification/implementation.
If your proteins are spread across multiple panels
- Provide the cross‑panel distribution and explain why each target matters.
- Clarify whether you need all targets in one readout or can split aims across sub‑studies.
- Flag sample volume constraints, time points, and any special matrices.
If you only know the biology, but not the exact targets yet
- Describe the biological decision and pathways of interest; share any discovery findings.
- State the study stage and sample design; outline matrix context and constraints.
- Ask for a pragmatic route (standard vs Flex vs Focus) based on feasibility and evidence.
Embedded mini‑cases:
- Case C (urgent mix): A biotech team requested "Explore + Focus + Signature Q100." We reframed the conversation to the biological aims and stage first; only then did the right combination become clear.
- Case A (running): The adipokine shortlist, two time points, and plasma context made it straightforward to check Flex library coverage before considering Focus.
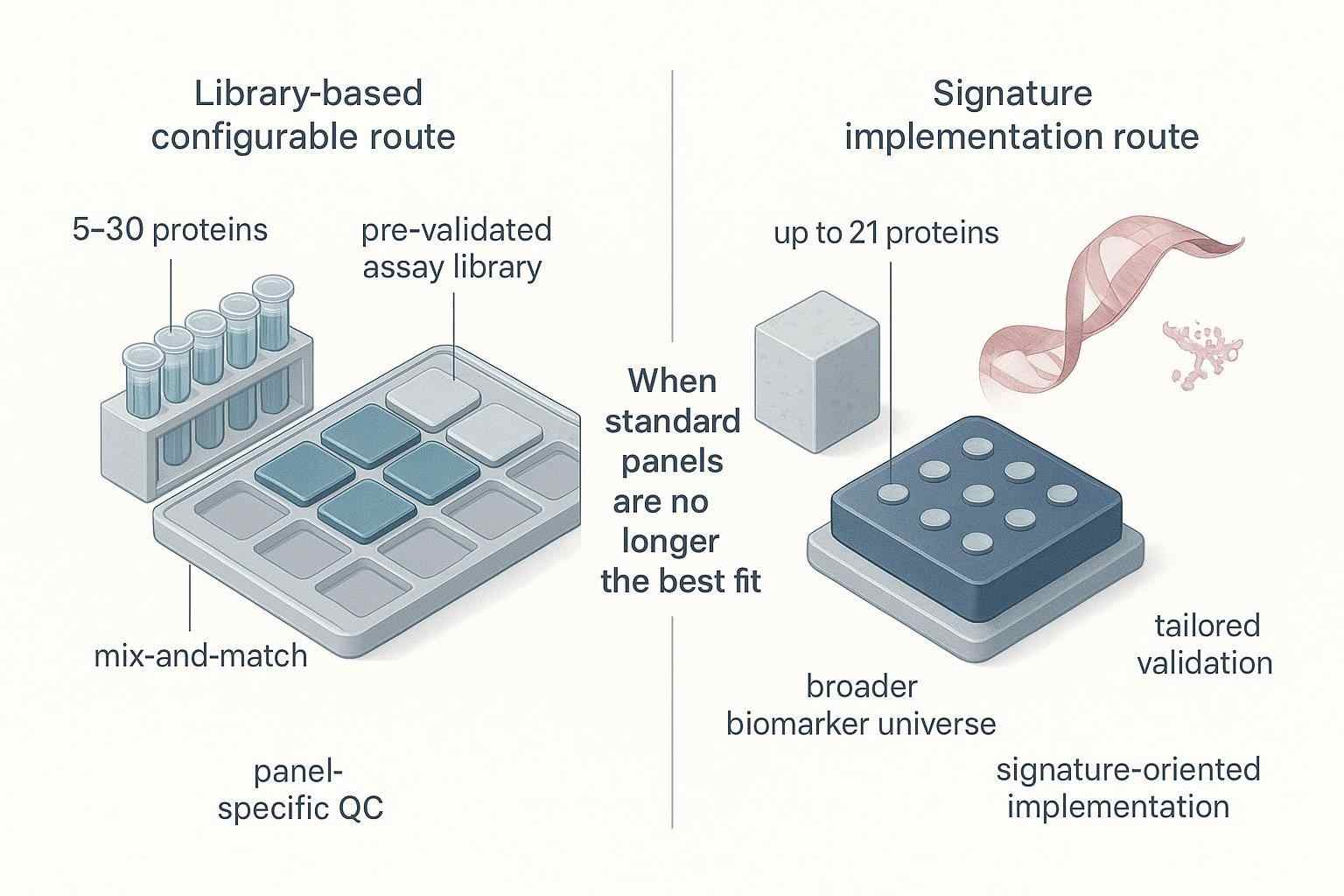 Figure 3. A well‑scoped custom biomarker request starts with protein priorities, study stage, matrix context, and sample structure—not just a product name.
Figure 3. A well‑scoped custom biomarker request starts with protein priorities, study stage, matrix context, and sample structure—not just a product name.
For broader context on discovery‑scale vs follow‑up trade‑offs, you may find these neutral primers useful: Olink High‑Throughput Proteomics and Olink vs Mass Spectrometry. When immune markers are in scope, these short reads can help teams articulate priorities: Olink Cytokine Panels and Immune System Research and Olink Cytokine Panel vs Other Methods.
As a neutral example of where a partner can help, Creative Proteomics supports scoping small custom sets, sample‑efficient designs (down to microliter‑scale matrices), and downstream bioinformatics for signature‑centric readouts (Knowledge Base Source).
Common mistakes researchers make when asking for a custom Olink panel
Confusing "small panel" with "simple project"
A 6‑protein panel may still demand careful validation, matrix checks, and QC planning. Small set ≠ trivial project.
Assuming every target can be combined instantly
Combinability is high in Flex, but not universal. Check library availability, assay compatibility, and validation needs before committing.
Requesting a custom panel before clarifying the study stage
Exploration, verification, and implementation imply different panel choices and metrics. State where you are and where you're going.
Asking for product names before defining the biological question
Lead with the decision you need to make and the proteins (or biology) that matter. Product selection follows from that logic.
FAQ
When is a standard Olink panel still the best choice?
When your biology aligns with an existing disease‑area panel and you still benefit from broad coverage for hypothesis generation, clustering, or pathway mapping. Standard panels offer coherent, validated content and can be the fastest way to produce interpretable results without over‑customizing prematurely. If you can't yet name a short list of must‑have proteins, stay standard.
What is the clearest sign that a standard panel is too broad for my study?
If only 5–10 proteins truly matter to your analysis or decision, and those targets are scattered across panels, a standard panel starts to look inefficient. This is especially true in verification or implementation stages, where interpretability and focus outweigh broad coverage.
Can I build a custom set for just 5–6 proteins of interest?
Often yes—first check whether all targets are present in the Flex library. If they are, a ~5–30 protein Flex panel can concentrate measurement on what matters most. If key targets aren't in the library or the aim is to implement a defined signature with tailored validation, consider Focus.
What should I do if my target proteins are spread across different Olink panels?
List the proteins, rationale, and matrix context. Determine whether you need a single readout or can split aims. If library coverage exists, a Flex set can avoid running multiple broad panels. If coverage is incomplete or validation needs are high, Focus may be the better route.
Is Olink Flex mainly for selecting known proteins from an existing library?
Yes. Flex is a semi‑custom path built around a pre‑validated assay library—typically ~5–30 markers from ~200 assays, with panel‑specific QC and high combinability (Olink Flex). It's efficient when your targets already exist and you want to avoid unnecessary breadth.
Is Olink Focus better suited to signature validation after discovery?
That's the common pattern. Focus supports up to 21 proteins with tailored development, drawing on a broader biomarker universe (>5,000) and aiming to implement or verify a defined signature (Olink Focus).
Do I need prior discovery data before considering Focus?
Typically yes—or at least a biologically defined, justified signature. Focus is about implementing what you already know you need to measure, with the validation depth and instrumentation to support that intent.
What information should I include when requesting a custom panel discussion?
Provide a prioritized protein list (or biology with candidate markers), species, matrix, sample counts and time points, study stage (exploratory vs verification/implementation), and any constraints around units (NPX vs pg/mL). If longitudinal or multicenter, add pre‑analytical considerations and QC expectations.
Conclusion
Standard panels are excellent when your study fits predefined biology and benefits from breadth. Flex is strong when you need a smaller, library‑based custom set—especially for cross‑panel target lists. Focus becomes the more logical choice when the project is about implementing or deeply validating a defined signature. The best starting point is not "Which product is cheapest?" but "What biological decision am I trying to make?" If you want a pragmatic conversation grounded in study stage and assay feasibility, you can start with a concise brief—and, where appropriate, explore neutral scoping support from Creative Proteomics (Knowledge Base Source).
Author: CAIMEI LI, Senior Scientist at Creative Proteomics
References (selected): According to Olink product documentation and peer‑reviewed literature
- Olink Flex product page: Olink Flex
- Olink Focus product page: Olink Focus
- Olink Signature Q100 instrument: Olink Signature Q100
- Peer‑reviewed: Carlyle 2022, Frontiers in Neurology; Heylen 2024; Siegbahn 2023

