If you work on Olink PEA studies long enough, you'll hear the same design questions again and again: "Can we run duplicates?" "Is one well enough?" "If our plasma volume is precious, should we sacrifice duplicates or preserve more subjects?" The truth is, many projects don't need blanket duplicate wells. What they need is clarity: What uncertainty are we trying to reduce—technical noise or biological variation—and what's the smartest way to spend limited sample volume?
Consider two real-world planning moments. Mini‑case A: a translational team wants to profile both clinical plasma and bone marrow plasma across four panels. They're debating whether duplicates are necessary on every sample or whether that would crowd out biological breadth. Mini‑case B: a fibroblast conditioned media experiment asks whether a single well per sample is sufficient, or if adding more technical repeats would be wiser than preparing more independent cultures.
Here's the deal: on Olink, internal/external controls, NPX‑based normalization, and plate randomization already address part of the technical question. Technical replicates can still be valuable—but as a design tool for specific goals (precision estimates, matrix de‑risking, or subset checks), not as a default courtesy. And they never replace biological replicates for inference.
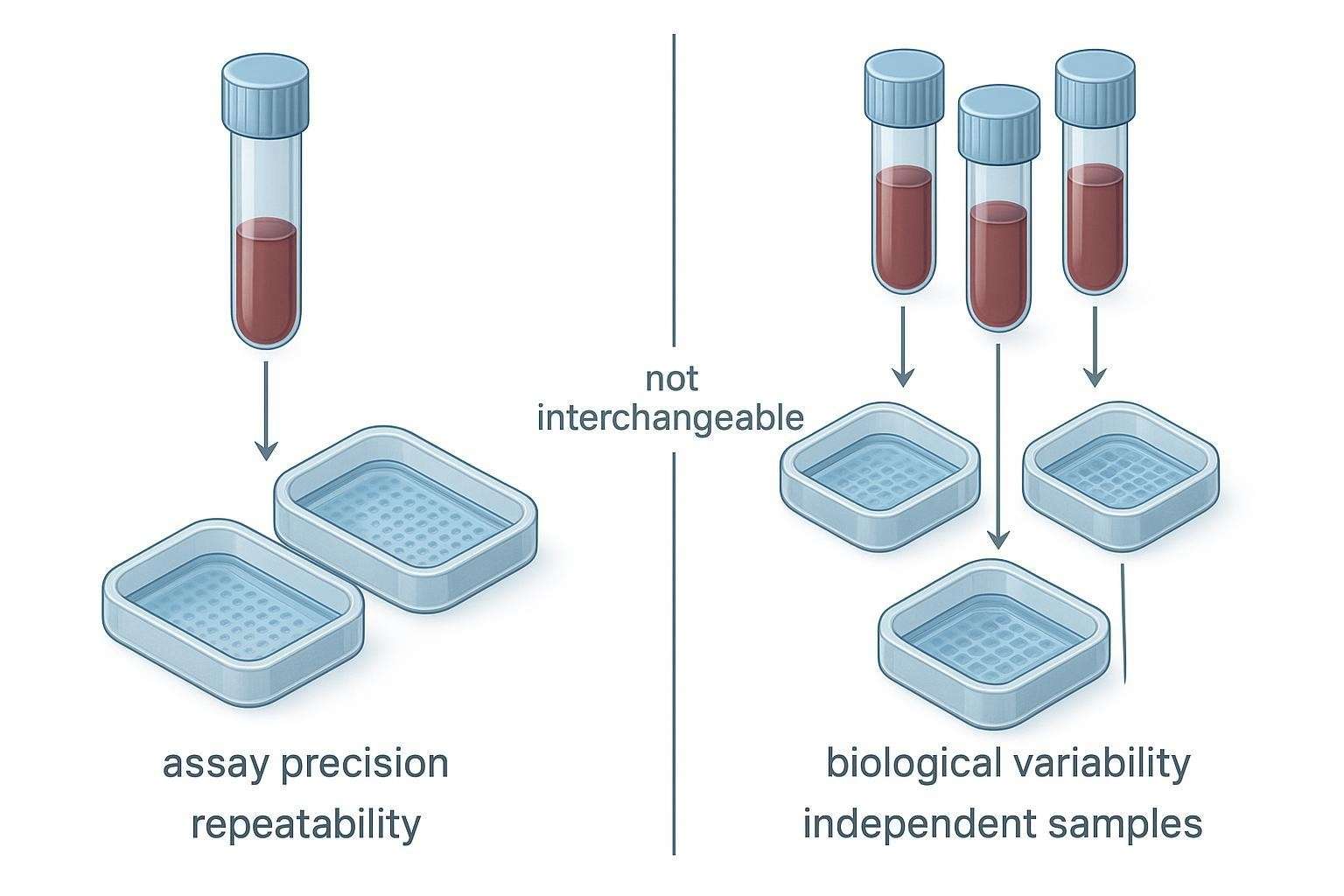 Figure 1. Technical and biological replicates reduce different kinds of uncertainty in Olink study design and should not be treated as interchangeable.
Figure 1. Technical and biological replicates reduce different kinds of uncertainty in Olink study design and should not be treated as interchangeable.
Technical replicates and biological replicates do not answer the same question
What technical replicates estimate
Technical replicates are repeat measurements of the same biological sample. They primarily estimate repeatability: the spread introduced by the assay workflow and measurement process within a narrow scope (e.g., intra‑plate variation). In Olink terms, duplicates can provide a more granular read on within‑plate precision in a pilot, or help confirm that a difficult matrix behaves acceptably under standard QC. They are useful when the study goal involves understanding technical spread or validating feasibility before scaling.
What biological replicates estimate
Biological replicates are independent samples—different subjects, animals, time‑matched independent culture preparations, or otherwise independent biological units. They capture biological variability and support inference about populations, groups, and effects. If your main question is, "Does group A differ from group B?" you'll get more inferential power by increasing biological N than by duplicating the same samples across wells.
Why confusing the two leads to weak conclusions
Treating technical replicates as if they were independent biological samples (pseudo‑replication) inflates N and can bias p‑values and confidence intervals. Rigor frameworks from NIH emphasize distinguishing the true experimental unit, being transparent about replication, and avoiding pseudo‑replication when reporting preclinical research; see the NIH's Principles for Rigor and Transparency and related guidance on replicates and independence in the Assay Guidance Manual. For background, consult the NIH's guidance on reproducibility and reporting and the Assay Guidance Manual's discussion of replicate types.
Technical vs biological replicates: what each one actually tells you
| Replicate type | What it measures | What it does not replace | Typical use in Olink planning |
| Technical replicates (duplicate wells of the same sample) | Repeatability/assay precision within a limited scope (often intra‑plate) | Independent biological sampling; they cannot stand in for additional subjects/animals/preparations | Pilot precision readouts; de‑risking non‑standard matrices; subset checks across plates |
| Biological replicates (independent samples) | Biological variability across subjects/animals/preparations | QC controls; they do not quantify technical repeatability by themselves | Powering group comparisons; cohort inference; time‑course/longitudinal analyses |
Why Olink projects often need better design decisions—not automatic duplicates
Internal controls, QC, and normalization already address part of the technical question
Olink integrates engineered internal controls per sample to monitor key steps of the PEA workflow and supports QC/normalization using NPX software. These platform features reduce the need to rely on blanket duplicates just to check that the assay is "working." For platform context, see Olink's overview of NPX software and normalization/standardization concepts in the Olink documents library. For a process‑level explanation of NPX, QC flags, and downstream analysis steps, you can also review the Creative Proteomics knowledge note on the Olink data analysis process.
- Olink NPX software overview: according to Olink's software pages, NPX calculation and QC workflows are standardized, leveraging internal controls at multiple steps.
- Normalization/standardization paper: Olink's knowledge "Documents" section describes how normalization methods and bridging support comparability without default duplicates.
- Internal resource: The Creative Proteomics knowledge base outlines how QC flags are interpreted and how NPX normalization is handled in practice.
Randomization and plate‑aware design often protect data quality more than duplicate reflexes
Balanced plate randomization protects against systematic bias. Olink's planning guidance emphasizes distributing cases/controls (or arms) evenly across plates and leveraging analytical tools for bridging where appropriate. When you randomize well and plan plates with awareness, you reduce the marginal benefit of running everything in duplicate simply to guard against plate effects. For planning pointers, see Olink's article on practical tips for multiplex immunoassay planning, which highlights randomization fundamentals.
Duplicates are a design choice, not a default courtesy
The most robust stance is not "always duplicate" or "never duplicate," but "use duplicates deliberately when they answer a specific design question." In practice, researchers often ask for duplicates before defining what uncertainty they are trying to reduce. Resist that impulse; frame duplicates as a targeted tool.
When Olink technical replicates are actually worth considering
Scenario 1: Pilot studies where you need a read on precision or feasibility
Early pilots—think mouse N=5 per group, or a first pass in a novel matrix—benefit from a small number of technical duplicates to estimate within‑plate precision and to check for obvious matrix‑related anomalies. The point is not to inflate N, but to quantify technical spread and decide whether to proceed, adapt panels, or invest in a larger cohort. In Mini‑case A, a team might reserve duplicates for a handful of bone marrow plasma samples during the first run to validate feasibility before committing the full panel set.
Scenario 2: Non‑standard or difficult matrices
Conditioned media, bone marrow plasma, certain ocular fluids, even DBS pilots—these can behave differently from conventional plasma/serum. Strategic duplicates on a representative subset can surface matrix‑specific effects (e.g., viscosity, protein background) early. As Olink's blog on challenging samples notes, matrices vary widely; rather than duplicating everything, consider duplicates where they most de‑risk interpretation.
Scenario 3: Projects where a subset of duplicate samples can de‑risk scale‑up
In mid‑stage projects, duplicating a small, representative subset per plate (or across plates) helps you confirm technical stability and supports bridging if the study expands over time. This subset strategy preserves biological breadth while still giving you plate‑aware precision checks. In Mini‑case B, instead of duplicating every conditioned media sample, you might duplicate a few independent preparations plus media blanks to verify that the workflow precision and background behave as expected.
Situations where technical replicates may add real value in Olink studies
| Project situation | Why duplicates may help | Better alternative if sample is limited | What to clarify before deciding |
| Small pilot/feasibility | Estimate intra‑plate precision; detect workflow outliers early | Keep duplicates to a subset and expand biological N if go/no‑go hinges on group effects | Primary question (precision vs inference), panel choice, planned plate count |
| Non‑standard matrix (e.g., conditioned media, bone marrow plasma, DBS) | De‑risk matrix behavior; check background and recovery | Representative subset plus appropriate blanks/controls; preserve main cohort | Known matrix risks, available volume per sample, replaceability |
| Scale‑up de‑risking (multi‑plate or multi‑batch) | Confirm stability across plates; aid bridging decisions | Use bridge/pooled QC samples; duplicate only a small per‑plate subset | Plate layout, bridging plans, whether variance is mainly technical or biological |
Mini‑scenarios from practice:
- A mouse disease‑vs‑control pilot (N=5/group) benefits more from adding animals than from duplicating every animal; keep a few duplicates for a per‑plate precision readout.
- A longitudinal design with limited volume should prioritize time‑point coverage and balanced randomization; reserve duplicates for a small subset if you need reassurance about technical spread.
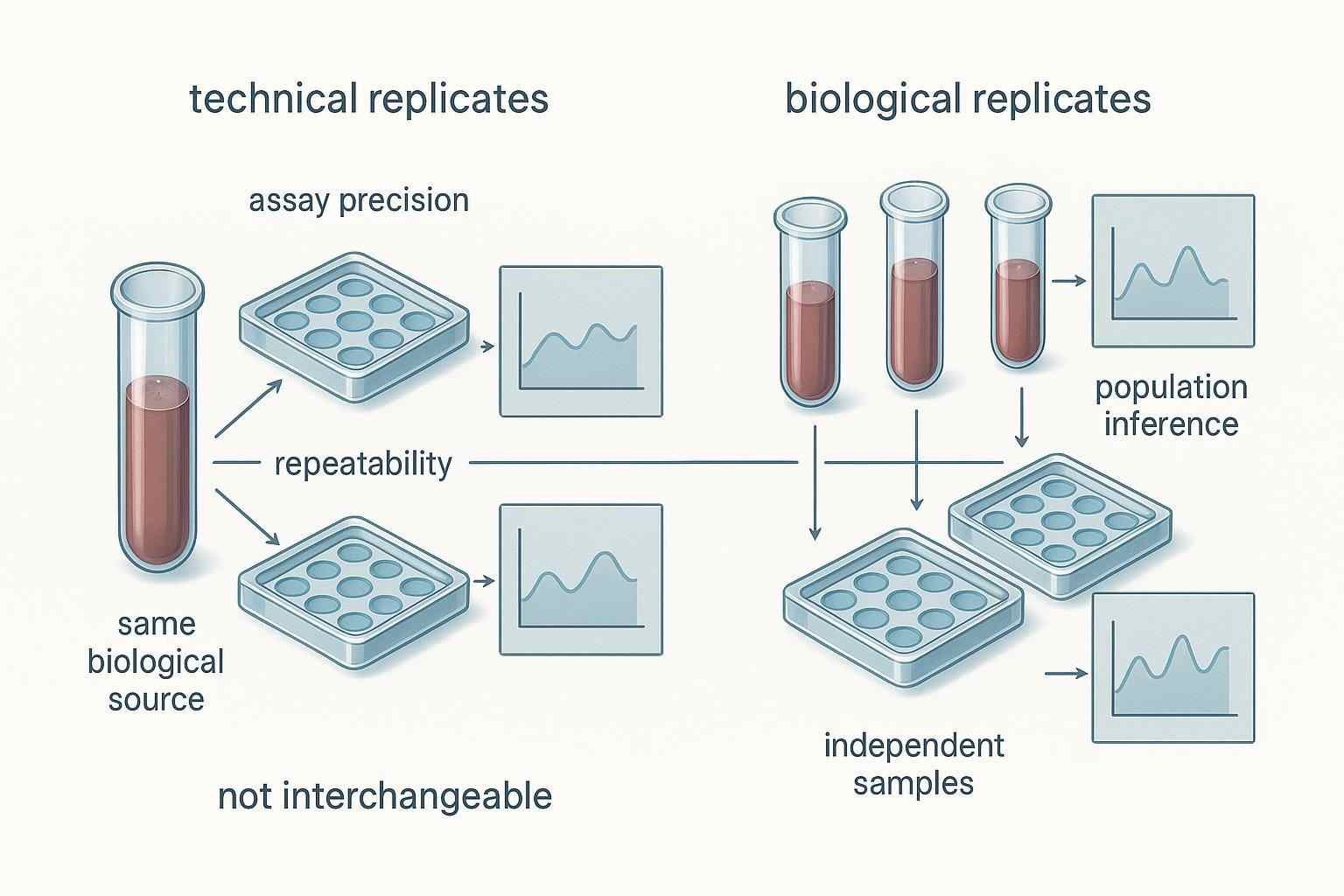 Figure 2. Technical replicates are most informative when they are used strategically—for pilot risk reduction, difficult matrices, or representative subset testing—not simply by default across all samples.
Figure 2. Technical replicates are most informative when they are used strategically—for pilot risk reduction, difficult matrices, or representative subset testing—not simply by default across all samples.
When more biological samples usually matter more than duplicate wells
Human cohort and translational studies
If your aim is to infer across patients—group differences, associations, stratification—then independent biological samples dominate value. In well‑randomized, plate‑aware designs that rely on Olink controls and NPX normalization, duplicating every subject rarely changes the scientific conclusion. Use duplicates selectively (e.g., subset or bridge‑like roles) while preserving biological breadth. For broader strategy trade‑offs (panel depth vs scale), see Creative Proteomics' note on high‑throughput Olink planning.
Small preclinical studies
Duplicating the same few animals does not create generalizability. If you plan to compare groups, prioritize more animals first; keep a few duplicates for precision estimation only if the matrix or workflow is unfamiliar. NIH's reproducibility guidance underscores identifying the true experimental unit and avoiding pseudo‑replication—principles that apply just as much to preclinical proteomics.
Longitudinal or multi‑time‑point designs
Time‑structured designs already consume volume and budget. Protect your time‑point coverage, randomize carefully across plates, and consider bridge samples where needed. Use subset duplicates only if you cannot otherwise address a specific technical uncertainty. If your question is about within‑subject change over time, technical duplicates won't substitute for consistent sampling and plate‑aware layout. When expanding beyond proteomics, consider how multi‑omics can strengthen inference without duplicating wells; see Creative Proteomics' overview on integrating Olink with other omics.
One well enough? Thinking about replicates in conditioned media and cell‑based experiments
Why this question comes up so often
Cell‑based systems and conditioned media are common in discovery and translational studies. Available volume can be tight; collecting independent culture preparations takes time. Teams therefore ask, "Is one well enough?" or "Should we add more wells from the same flask?" The right answer depends on whether your added wells are truly independent biological units or just more measurements of the same preparation.
When "another well" is technical, not biological
If multiple wells come from the same culture preparation at the same time, they are very often technical replicates: repeated measurements of one biological unit. Treating them as independent samples risks pseudo‑replication. Rigor resources from NIH and the Assay Guidance Manual stress distinguishing technical repeats from biological replication and counting the true experimental unit correctly. In Mini‑case B, extra wells drawn from the same fibroblast preparation mostly narrow technical uncertainty; they do not broaden the biological claim.
What matters more than simply adding duplicate wells
For conditioned media and co‑culture supernatants, prioritize independent culture preparations (separate passages or separate flasks), include media/background controls, and map your total available volume. If you still want a read on technical spread, keep duplicates to a small subset selected for matrix risk. In our Mini‑case B, a sensible plan was: three independent fibroblast preparations as the biological N, media blanks, and duplicate wells for only one or two preparations to confirm precision—preserving the rest of the volume for biological breadth.
How to plan duplicates without wasting precious samples
If volume is limited
Start by stating the study question: precision estimation versus group inference. Allocate volume to independent samples first, then reserve a small fraction for subset duplicates if you need reassurance. Consider panel selection and plate count trade‑offs alongside replicate allocation; if a narrower panel frees volume for more subjects, that may serve your primary endpoint better.
If the sample cannot be recollected
When recollection is unlikely (e.g., rare clinical samples, limited bone marrow plasma), treat duplicates as an allocation decision. Use representative subset duplicates on the riskiest matrix types, and place bridge/pooled QC samples to enable plate‑aware checks. Protect the main cohort's biological breadth.
If you still want some measure of technical reassurance
Use a subset strategy: duplicate a small, plate‑balanced set (for example, one or two samples per plate, chosen to represent critical matrices or concentration ranges). Combine this with platform QC and NPX normalization to triangulate precision without consuming breadth. As an example of how subset‑duplicate matrix checks are documented in practice, see the Creative Proteomics knowledge note on the Olink data analysis process, which explains where QC flags and normalization fit into downstream interpretation.
A practical decision framework: should your Olink project include duplicates?
Start with the study question
Write down the primary contrast and what success looks like. If the answer depends on population‑level inference, push biological N. If the answer depends on feasibility or a precision threshold for go/no‑go, consider a small number of technical duplicates in a pilot or subset.
Then assess matrix risk and sample replaceability
Rank matrices by risk (e.g., bone marrow plasma higher than standard plasma; conditioned media with additives higher than simple media). Note which samples can be recollected and how much volume you truly have. High‑risk, irreplaceable matrices are where subset duplicates do the most work per microliter.
Finally decide whether duplicates belong in all samples, a subset, or only a pilot
Default to "no blanket duplicates." If your main uncertainty is technical and matrix risk is high, use subset duplicates. Reserve broader duplicate use for narrowly defined, precision‑driven validation studies where biological breadth is already sufficient.
Decision framework: no duplicates, subset duplicates, or broader biological expansion?
| Decision factor | Favor more biological samples | Favor subset duplicates | Favor broader duplicate use |
| Primary goal | Population/group inference; discovery breadth | Matrix/feasibility de‑risking before scale‑up | Precision validation as the endpoint |
| Matrix risk | Standard plasma/serum with low risk | Non‑standard matrices (bone marrow plasma, conditioned media, DBS) | Confirmed matrix quirks needing tight precision monitoring |
| Replaceability/volume | Readily recollectable; adequate volume | Limited volume; partially replaceable | Irreplaceable samples where precision itself is the deliverable |
| Plate/batch plan | Strong randomization; bridging strategy in place | Need to verify stability across selected plates | Formal method validation across plates/batches |
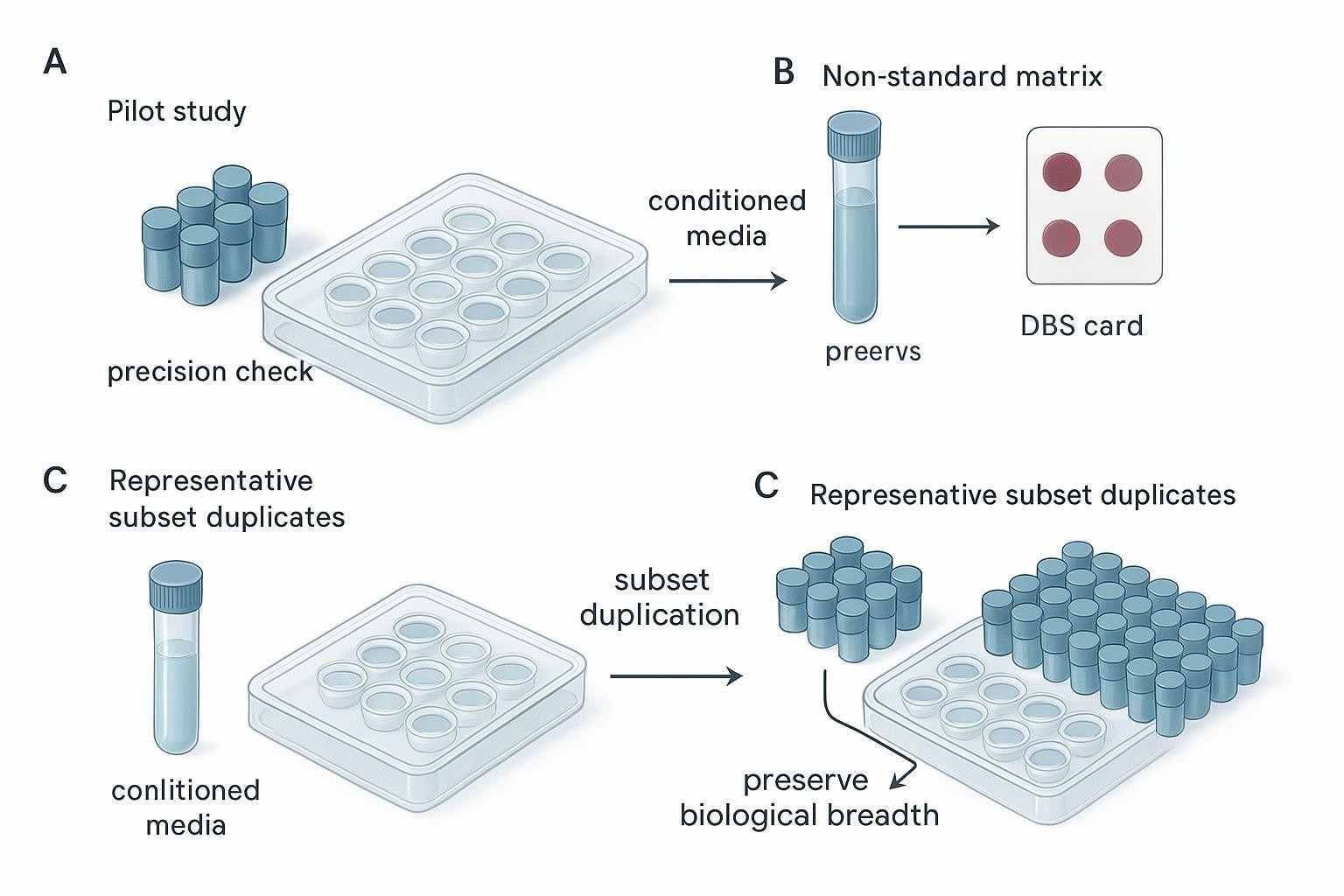 Figure 3. In Olink study planning, the best replicate strategy depends on the study question, matrix risk, sample replaceability, and whether the main uncertainty is technical or biological.
Figure 3. In Olink study planning, the best replicate strategy depends on the study question, matrix risk, sample replaceability, and whether the main uncertainty is technical or biological.
What to mention in your first inquiry if you are unsure about replicates
For cohort studies
State the primary contrast (e.g., case vs control), anticipated sample size, and plate count. Confirm whether you plan panel combinations or a single panel. Note any plan for bridging across time. Share your initial view on duplicates (none vs subset) and the trade‑off you worry about.
For non‑standard matrices
Describe the matrix (e.g., bone marrow plasma, conditioned media) and any known risks (hemolysis, viscosity, media additives). Include available volume per sample and whether recollection is possible. Ask about subset duplicates targeted to matrix risk. For preanalytics, see Creative Proteomics' Olink sample preparation guidelines.
For cell culture or limited‑volume projects
Specify whether additional wells would be independent preparations or technical repeats from the same culture. Provide total available volume and planned controls (e.g., media blanks). If panel selection is still open, consider whether a 96‑ vs 48‑plex choice affects replicate allocation; see this overview of Olink panel trade‑offs.
FAQ
Do Olink studies always need technical replicates?
No. Olink integrates internal controls and standardized NPX workflows that already address part of the technical question, and good plate randomization further reduces bias. Technical duplicates are useful when they answer a specific design goal—such as estimating within‑plate precision in a pilot or de‑risking a difficult matrix—not as a universal rule. For randomization context, see Olink's planning tips article; for normalization background, see Olink's documents on NPX and standardization.
What is the difference between a technical replicate and a biological replicate?
Technical replicates repeat the measurement of the same sample to estimate assay repeatability; biological replicates are independent samples that capture biological variability. Only biological replicates support population‑level inference. Treating technical repeats as independent samples is pseudo‑replication, a practice cautioned against in NIH rigor guidance and the Assay Guidance Manual.
If my clinical samples are limited, should I prioritize more subjects or duplicate wells?
If your objective is inference across patients, prioritize independent subjects and use subset duplicates only where they reduce genuine technical uncertainty (e.g., on higher‑risk matrices such as bone marrow plasma). Protect biological breadth first, then reserve a small fraction of volume for representative subset checks. For matrix planning, see Creative Proteomics' sample preparation guidance.
Can duplicates help with non‑standard matrices such as conditioned media or bone marrow plasma?
Yes—strategic duplicates on a representative subset can reveal matrix‑specific behavior early, helping you decide whether to proceed or adjust panels. Pair subset duplicates with media/background controls and plate‑aware randomization rather than duplicating everything. Olink's discussion of challenging samples underscores that matrices vary, so a targeted approach is sensible.
Is one well enough for conditioned media cytokine profiling?
Often, yes—if you are primarily seeking a directional read, one well per independent preparation can be sufficient, given Olink's QC and NPX normalization. If you need reassurance about precision, consider duplicating only a small subset (e.g., one or two independent preparations) and include media blanks. For interpretation pitfalls in serum/plasma studies that also apply conceptually, see this Creative Proteomics note on interpreting Olink serum proteomics.
Should I run all samples in duplicate, or only a subset?
Default to subset or pilot duplicates unless your study's primary endpoint is precision validation. Choose a small, plate‑balanced subset representative of matrices and concentration ranges. Maintain strong randomization and, where relevant, use bridge/pooled QC to monitor plate effects across time.
Do small mouse studies benefit from technical replicates?
They benefit most from more animals. A few technical duplicates can quantify intra‑plate precision in a pilot, but they don't substitute for independent biological replicates. Align choices with NIH principles on identifying the experimental unit and avoiding pseudo‑replication.
How should I describe my replicate concerns when requesting an Olink study?
Share your study question, matrix type(s), available volume, replaceability, plate count, and whether you are considering no duplicates, subset duplicates, or broader duplication. Note the trade‑off you fear (e.g., duplicates might crowd out biological breadth). This information allows rapid scoping and tailored design advice.
Conclusion
Technical duplicates can be genuinely useful in Olink studies—especially for pilot precision checks, matrix de‑risking, or subset validations—but they are not a universal requirement. In many translational and cohort settings, biological replicates, careful randomization, platform controls, and NPX‑based normalization matter more than automatic duplicate wells. For precious or non‑standard matrices, a subset‑based or pilot‑based strategy often delivers the best trade‑off. The right answer starts with the study question, not with habit. If you'd like preliminary design input, submit a brief "replicate expectation" template (study question, matrices, volume, replaceability, and current view on duplicates), and we'll share practical options for your scenario.
Author: CAIMEI LI — Senior Scientist at Creative Proteomics
References and resources
- Olink NPX software overview and QC workflows: https://olink.com/software/npx-software
- Olink documents: Data normalization and standardization: https://olink.com/knowledge/documents
- Olink Analyze (R) tutorials, including bridging guidance: https://olink.com/software/olink-analyze
- Olink planning tips with randomization guidance: https://olink.com/blog/practical-tips-for-planning-a-successful-multiplex-immunoassay-experiment
- Olink Flex page (internal controls context): https://olink.com/products/olink-flex
- Olink blog on challenging samples: https://olink.com/blog/working-with-challenging-samples-for-additional-layers-of-biological-insights
- NIH Principles and Guidelines for Reporting Preclinical Research: https://grants.nih.gov/policy-and-compliance/policy-topics/reproducibility/principles-guidelines-reporting-preclinical-research
- Assay Guidance Manual (replicates and independence): https://www.ncbi.nlm.nih.gov/books/NBK550206/
Internal Creative Proteomics knowledge links
- Olink data analysis process (NPX/QC overview): /knowledge/olink-data-analysis-process.html
- Olink sample preparation guidelines: /knowledge/olink-sample-preparation-guidelines.html
- High‑throughput Olink & scaling: /knowledge/olink-high-throughput-proteomics.html
- Interpreting Olink serum proteomics: /knowledge/interpreting-olink-serum-proteomics.html
- 96‑ vs 48‑plex panel trade‑offs: /knowledge/olink-96-vs-48-plex-panels.html
- Integrating Olink with other omics: /knowledge/integrating-olink-with-other-omics.html

